The first DNA programmable computer
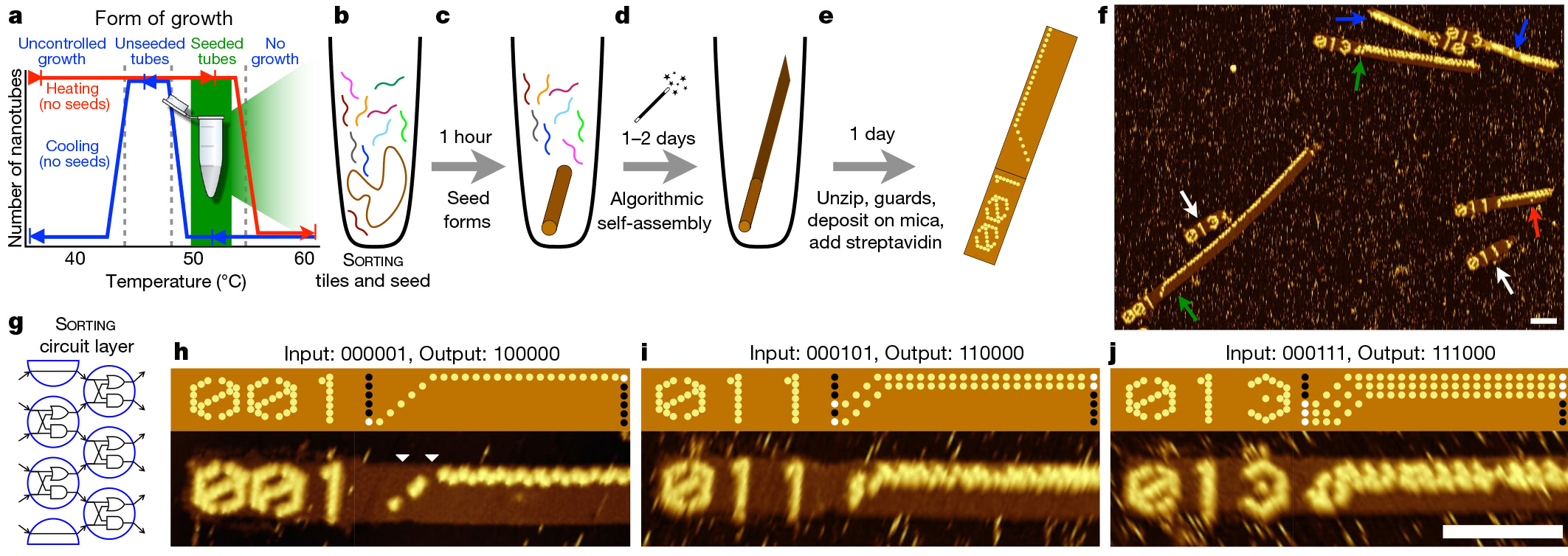
Experimental protocol and implementation of the sorting algorithm on a programmable DNA computer
Scientists have long been experimenting with storing information in DNA and processing this information. For example, scientists from Washington University and Microsoft recently built the "world's first DNA Winchester" ( photo ). This design is able for the first time to provide the recording and reading of information in DNA storage without human intervention. A very significant achievement, given that DNA can record information with a density of 2.2 petabytes per gram . DNA is a compact container with a recording density thousands of times larger than existing carriers.
However, all existing DNA systems have a problem: all these are unique proprietary developments that completely lack any flexibility. If we compare with silicon technology, then each group of researchers from scratch develops a new computer architecture, for which you need to write new software. But things can change thanks to the first programmable DNA computer developed at the University of California, Davis (UC Davis), the California Institute of Technology, and the University of Maynooth.
The first DNA programmable computer is described in a scientific article., which was published on March 20, 2019 in the journal Nature. The authors showed that with the help of a simple trigger, the same basic set of DNA molecules is able to implement many different algorithms. Although the study is a purely laboratory experiment, programmable molecular algorithms in the future can be used, for example, to program DNA robots that already successfully deliver drugs to cancer cells .
“This is one of the landmark works in this area,” says Torsten-Lars Schmidt, an assistant professor of experimental biophysics at the University of Kent, who was not involved in the research. “They used to demonstrate algorithmic self-assembly, but not to that degree of complexity.”
In electronic computers, bits are binary units of information. They represent the discrete physical state of the basic equipment, for example, the presence or absence of an electric current. These bits, or rather electrical signals, are passed through circuits consisting of logic elements that perform an operation on one or more input bits and produce one bit as an output.
Combining these simple building blocks over and over, computers can run amazingly complex programs. The idea of DNA computing is to replace electrical signals with chemical bonds, and silicon with nucleic acids to create biomolecular software.
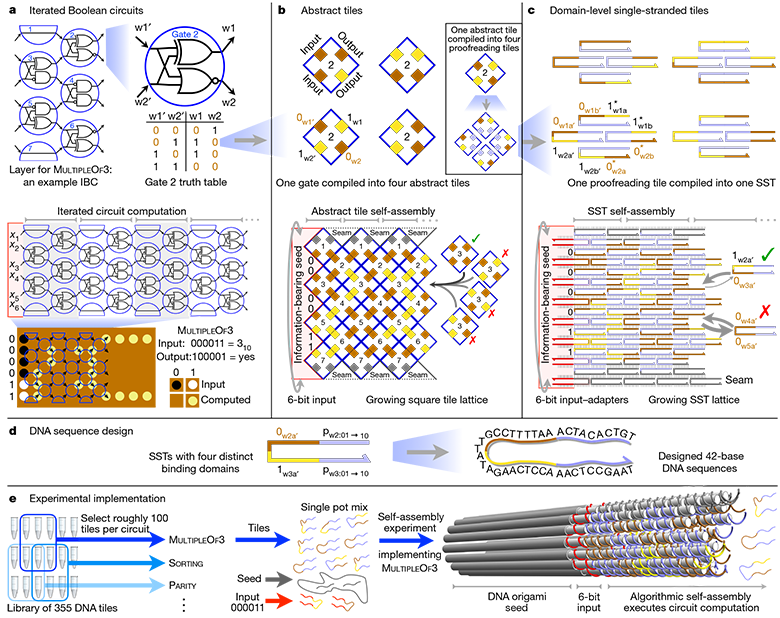
Abstract hierarchy of architecture and practical implementation of the full 6-bit IBC (Iterated Boolean Circuit)
According to Eric Winfrey, a scientist at the California Institute of Technology and co-author of the article, molecular algorithms use the natural possibilities of processing information in DNA, but instead of allowing nature to take reins of government, calculations in DNA are made in accordance with a program written by man.
Over the past 20 years, several successful experiments with molecular algorithms have been carried out, for example, for playing tic-tac-toe or assembling molecules of various shapes. In each case, careful development of the DNA sequence was required in order to execute one specific algorithm that would generate the DNA structure. In this case, the difference is that the researchers developed a system in which the same basic DNA fragments can be ordered to create completely different algorithms - and, therefore, to obtain completely different results.
The process begins with the DNA origami technique, that is, folding a long chain of DNA into the desired shape. This folded slice works like a seed, which runs an algorithmic assembly line. Seed remains virtually unchanged, regardless of the algorithm. For each experiment, only small changes are made to it in several sequences. Reprogramming the logic circuit
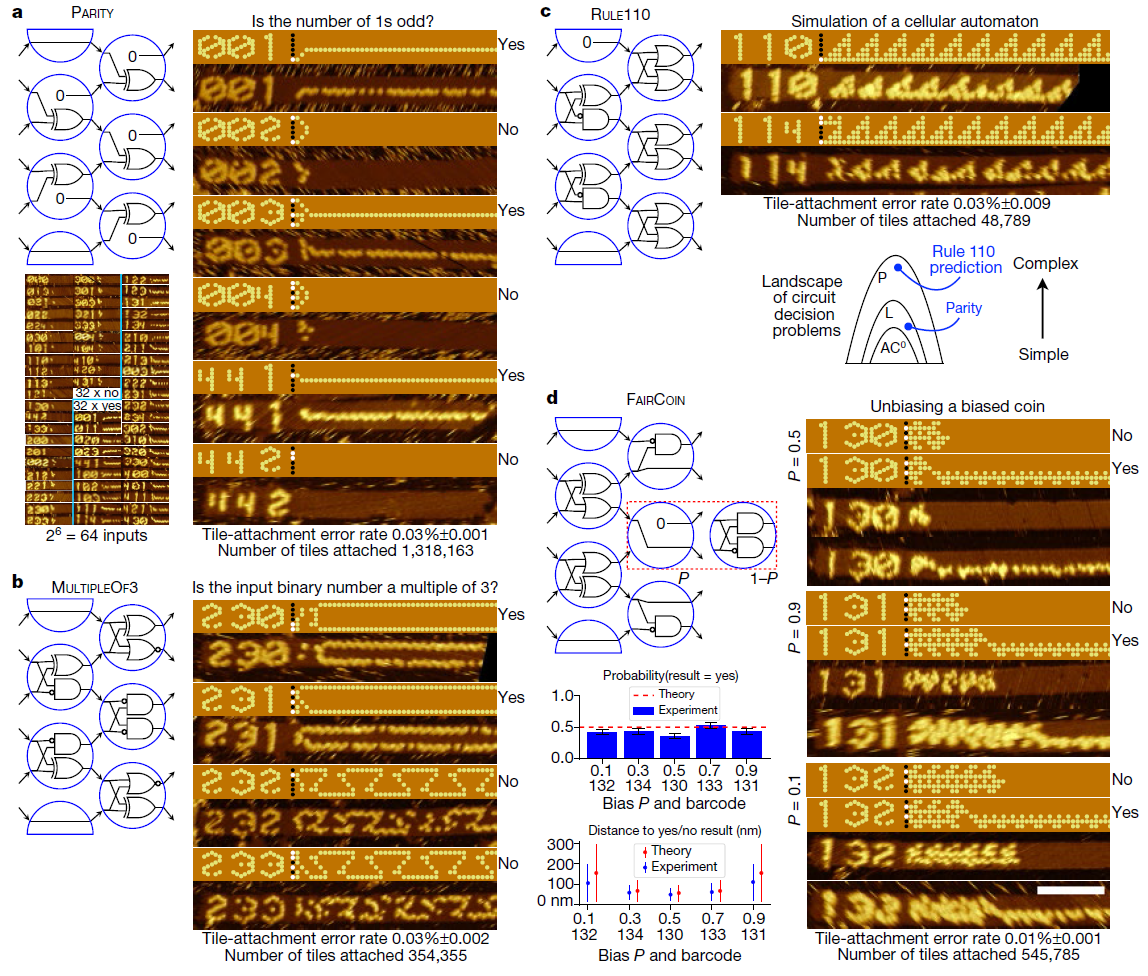
After creating the “seed”, it is added to the solution with hundreds of other DNA strands, known as DNA tiles. Scientists have developed 355 of these tiles. Each has a unique arrangement of nitrogenous bases. Accordingly, for each algorithm, researchers simply choose a different set of starting tiles. Since these DNA fragments are joined during the assembly process, they form a circuit that implements the selected molecular algorithm on the input bits provided by the “seed”.
Using this system, the researchers developed and tested 21 algorithms for performing tasks such as recognizing a division by three, choosing a leader , generating patterns, and counting from 0 to 63. All these algorithms are implemented using different combinations of the same 355 DNA tiles.
Of course, it is not easy to write code by dropping DNA fragments into a test tube, but if the process is automated, future molecular programmers do not even have to think about biomechanics, as today's programmers do not need to understand the physics of transistors in order to write good programs.
