Tasks of domestic sequencing (June, 2019)
Before reading this upgrade, it is advisable to familiarize yourself with the source code published earlier .
The very first and most popular nanion pore sequencer MinION (the minimum set in the UK is $ 1000, in Russia - 150 thousand rubles), developed by Oxford Nanopore Technologies (ONT), operates on disposable cells, each of which costs $ 900 (in Russia - 135 thousand rubles). Such cells allow you to digitize DNA 10 ... 20 Gb in 2 ... 3 days. This is too little for sequencing the human genome (you need> 100 Gb), but too much for all other clinical diagnostic purposes (<1 Gb is enough).
Recently started selling more productive (and more expensive) models - PromethION 24/48 ($ 165,000 / $ 285,000), aimed primarily at full-genome in-line sequencing. More expensive are their more efficient consumables ($ 2,000 / unit,> 100 Gb). True, they can be cheaper in bulk ($ 1,800,000 for 2,880 pcs., $ 625 / pcs.), But such acquisitions (200 ... 250 million rubles per batch) are not permissible for the Russian budget.
And not only for Russian. Therefore, the most attention of visitors to the recent ONT conference (May 22-24) was attracted by the announcement of the start of sales of Flongle, an adapter insert for the MinION sequencer, which allows working with less efficient (~ 1 Gb), but relatively cheap ($ 90) disposable cells.

https://nanoporetech.com/products
The appearance of Flongle can cause an explosive growth in the use of NGS (next generation sequencing) in clinical practice and displace conventional PCR diagnostics, whose market size in Russia exceeds one billion rubles, and in the world it is measured in billions of dollars. But for most Russian patients, NGS diagnostics will still remain an inadmissible luxury, since the cost of cells for Flongle in the domestic market will be more than 10 thousand rubles. And if we add to it the cost of expendable reagents, transport costs and overhead, then the estimated cost of sequencing one DNA / RNA sample will not fall below 20 thousand rubles.
This implies the need to develop domestic nanopore sequencers and provide them with cheap consumables and reagents.
Significantly reduce the cost of nanopore sequencing is possible by reusing cells for Flongle and import substitution of the necessary reagents for working with them. But first you need to get a test device that allows you to monitor the formation of bilayer lipid membranes (BLM) in 126 wells of the working cell, and evaluate the quantity and quality of nanopores embedded in these membranes. Such a device ("Nanorider"?) Should track changes in the electrical conductivity (and / or impedance) of individual holes during the formation of BLMs on them, and register picoamperous currents flowing through single ion channels.
It is possible to compose a reader connected to the computer via USB 2.0 from standard elements: FPGA (100 ... 200 $), current amplifiers (MAX9923FEUB - 200 ... 300 rubles), ADC (1 ... 2 thousand rubles), etc. ... It’s necessary to develop only a board with a non-standard contact pad (10x13) type LGA (Land Grid Array) for docking with a cell from Flongle. Or with its simplified domestic counterpart containing a smaller number (16 ... 32) of sensor wells.

Touch Pad Cell Flongle
Reducing the design productivity of cells by 4 ... 8 times (up to 10 ... 20 Mb) is not critical for solving many problems (determining pathogens, HLA typing, establishing paternity, identification, etc.), but it allows you to collect a nanopore sequencer on conventional amplifiers picoampere currents. And do without the use of special 512-channel chips, the development of which the Oxfordians have invested tens of millions of dollars. True, each multichannel nanoreader must contain more than a dozen current amplifiers + ADCs, but it can still cost several times less than the proprietary MinION + Flongle combination (> 200 thousand rubles). As for the low performance of the nanoreader, for Russian users who do not have decent computers (doctors, biologists, biohackers, etc.), it can even be useful.
The 512-channel picoampere current amplifiers developed by the Oxfordians are contained in each disposable cell for the MinION. Waste cells can be thrown away, but it is better to use the chips contained in them to produce Flongle analogues that are characterized by increased productivity. If these chips are equipped with surface contacts, they will be able to work with contact cells containing 2048 (512x4) sensor wells.
Designing cells with such LGAs will not be easy, but you need to take into account that the contact requirements here are less stringent than processors and motherboards, in which one bad contact can disable the entire system. For a sequencer, the inoperability of even half of the cell contacts can be considered acceptable. The remaining half is enough to ensure performance at the level of 5 ... 10 Gb.
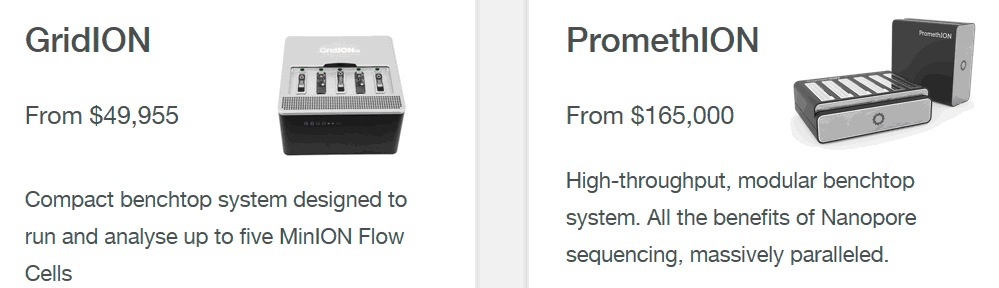
The logical development of this idea will be the combination of several similar readers in one device. This will allow you to get an analog of the GridION X5 sequencer, designed for simultaneous work with five (or more) reusable cells of domestic production. Such a sequencer may be in demand in many clinical diagnostic laboratories.

https://store.nanoporetech.com/devices
Disposable PromethION sequencer cells contain 9,000 sensor wells on chips. And the chips themselves are 3000-channel amplifiers of picoampere currents. The stream of information that they give out with 24 (PromethION 24) or 48 (PromethION 48) information cells is so large that in a raw form it cannot be transferred to a regular external computer. Therefore, sequencers of this type must be equipped with their own supercomputer. But if the sequencer will work only with one cell, then it can be connected to a regular gaming laptop with a Thunderbolt 3.0 port.
Alteration of free (ejected) chips from ONT to work with contact cells is not an insoluble problem. But sequencers built on the use of such chips can only work with ONT software. And the efforts that will have to be spent on remaking these programs can negate the benefits of using off-the-shelf chips. And developing such chips from scratch is too expensive.
This year, beta testing of the nanopore sequencer from AXBIO (USA) is due to begin. It uses chips with a million channels of current amplification produced in Japan by the Israeli company Tower Semiconductor Ltd ... Consumables and the sale of these sequencers will be handled by the Chinese branch of AXBIO.

In the United States and the European Union, more than 7,000 patents are devoted to nanopore sequencing, which creates unpredictable financial risks for local companies trying to deal with such technologies. Apparently, this is the main reason for such a high internationalization of this development. And it allows Russian companies and laboratories to participate without any restrictions in testing and improving AXBIO technology, as well as producing (or collecting) such sequencers on their territory.
Terabytes of raw data obtained by a nanopore sequencer with megapixel chips (mega reader), it is desirable to filter. And transfer to the computer only the longest and highest-quality reads. This will not only improve the quality of sequencing, but also reduce the requirements for the computer, which is also important for mass production and widespread use of genomic sequencers. In addition, the excessive productivity of mega readers can reduce the duration of sequencing of the human genome from 2 ... 3 days (for ONT) to several hours.
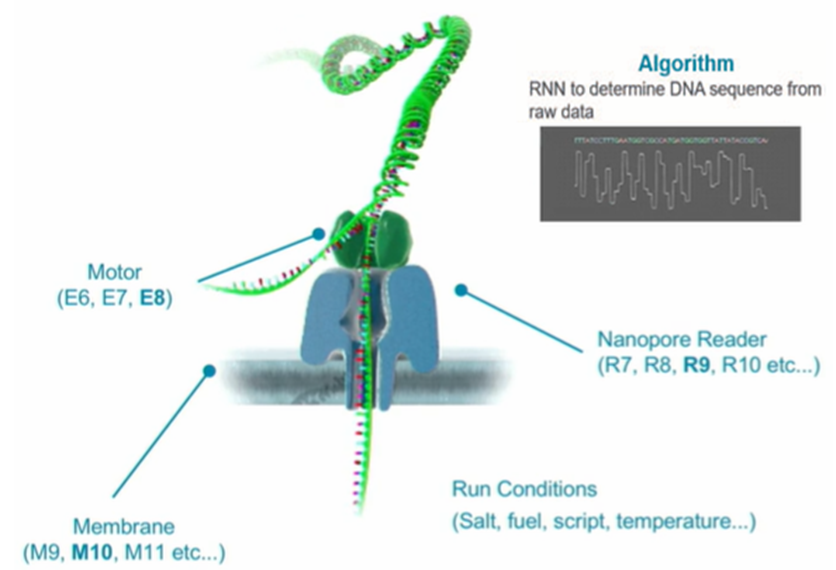
ONT's success was facilitated by the organization of parallel pipeline optimization of the five main elements of nanopore sequencing technology — membranes, nanopores, “motor” (helicase), sequencing conditions, and decoding algorithms for read signals.
https://nanoporetech.com/resource-centre/videos/sub1000
The guaranteed shelf life of cells for nanopore sequencing depends mainly on the stability of the bilayer lipid membrane. Now it is 6 weeks, although usually such membranes are destroyed in a few days. To increase their shelf life, hydrophobic polymer additives, surface-active block copolymers, cholesterol, antioxidants, cross-linking of fatty acids, etc. are used, and such modifications can interfere with the formation of ion channels. Therefore, it is very difficult to achieve BLM stability.
Radically solving this problem allows the formation of membranes in the working cells of the sequencer and the introduction of nanopores into them immediately before use. Self-preparation of cells for work will provide the opportunity for their regeneration and reuse. As for the shelf life, for dry cells it is almost unlimited, and the reagents necessary for their activation and regeneration can be stored for months.
The bottleneck in the technology of nanowire sequencing is at the same time the bottleneck of the nanopore, on which the quality of the read information depends. Therefore, one of the main tasks of ONT was the search for a membrane protein, which allows obtaining results of acceptable quality.
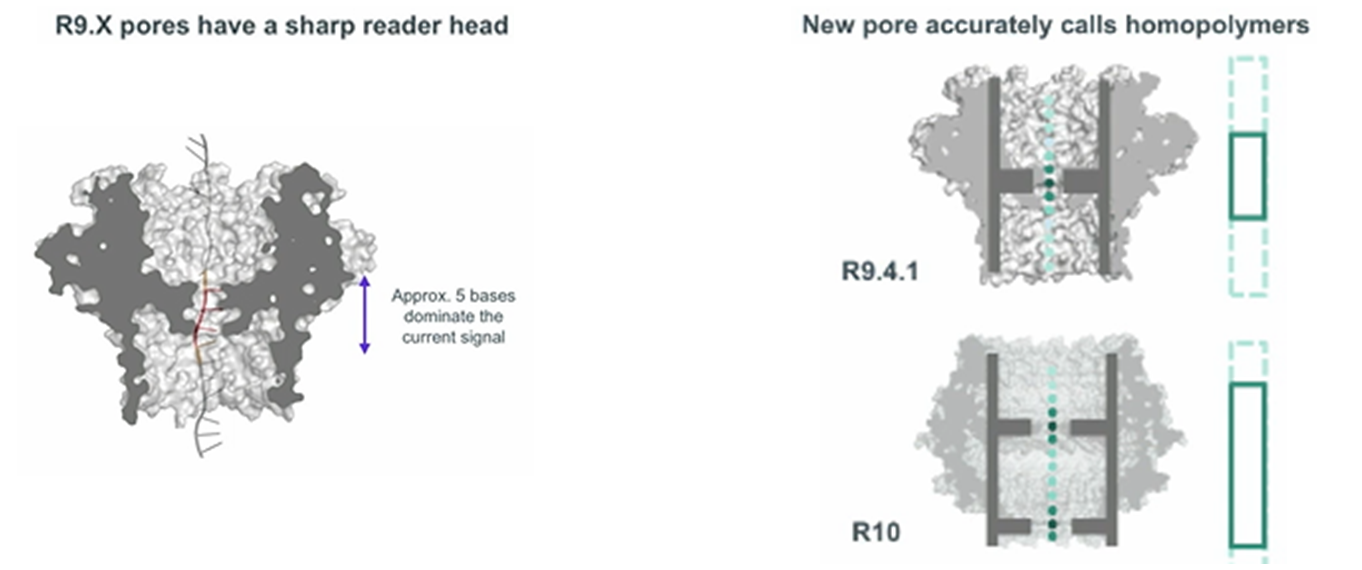
The first such protein was CsgG, an outer membrane protein found in many gram-negative bacteria. Nine of its subunits form an ion channel in the membrane with wide entrance and exit vestibules, and one bottleneck. But it was possible to obtain sufficiently high-quality nanopores (R9, R9.4.1, R9.5.1) only after enumerating and analyzing hundreds of modified CsgG variants. This summer, cells with the next generation pores (R10 or R10b) should appear on sale, featuring increased accuracy in reading homopolymer repeats.
https://nanoporetech.com/about-us/news/london-calling-clive-brown-and-team-plenary
ONT is an essential component of ONT's nanopore sequencing technology, a helicase, a molecular motor that unwinds double-stranded DNA and inhibits the progress of single-stranded DNA through a nanopore. The maximum speed of DNA unwinding with such a motor now reaches 450 nucleotides per second, although the speed of MinION electronic components allows it to be increased to 1000 nucleotides per second. Theoretically, accelerating the work of helicase can double the performance of MinION, but in practice this can lead to poor sequencing. Therefore, faster and more efficient helicases are unlikely to improve the performance of nanopore sequencers such as MinION and PromethION.
The quality of sequencing depends on many conditions - the magnitude of the voltage applied to the membrane, the electrical conductivity and the ionic composition of the reaction mixture, the amount and nature of impurities clogging the pores, the ability to clean clogged pores by changing the polarity of the potential, the structure of the adapters used, etc., etc. ... Therefore, nanopore sequencing requires careful optimization of all its parameters and compositions of working solutions (Run Conditions), which can adversely affect the quality of digitization of nucleotide sequences awns. For example, it was recently shown that a decrease in the concentration of ATP cleaved by helicase gradually decreases the accuracy of sequencing results, and now Oxfordians are trying to eliminate this problem by introducing an ATP regeneration system into the reaction mixture.
The fifth element of nanopore sequencing technology is artificial intelligence built on recurrent neural networks (RNN) and on the use of some special algorithms and programs that improve the recognition of homogeneous repeats (Flip-flop, Medaka). In July, the next release of the Guppi and MinKNOW programs should appear, in which the Oxfordians promise to add the ability to determine methylated bases (5mC and 6mA). This is important not only for epigenomic studies, but also to improve the recognition of conventional bases.
The large volumes of information read by nanopore sequencers and the complexity of its processing impose increased requirements on the hardware of such software and hardware systems. Therefore, not every computer is suitable for connecting MinION, the latest generation of graphics accelerators are installed in GridION and PromethION, and Oxford Nanopore Technologies became one of the first buyers of a large batch of Jetson AGX Xavier (NVIDIA) modules designed for devices with artificial intelligence.
More recently, nanopore technology was considered to be nothing more than a useful complement to more accurate fluorescence and semiconductor sequencing technologies. But a gradual increase in the accuracy of reading DNA, combined with a large length of readable sequences, allowed it, if not to get ahead of all competitors, then at least determine the main vector of development of genomic sequencing technologies.

It may take quite a while to master all the elements of nanopore sequencing technology, but this is not the biggest problem. The main problem is the lack of targeted funding for such integrated developments in Russia. True, the recently approved Federal Scientific and Technical Program for the Development of Genetic Technologies for 2019–2027 (section "Program Implementation Directions") says that in the short term (3 ... 6 years) a prototype device for high-performance genomic sequencing will be developed. But finding this single line on 30 pages of text is not so easy. Easier to negotiate with the Chinese on the purchase of AXBIO sequencers.
Even simpler is to find craftsmen who can assemble the simplest nanorider “on the knee”. And then walk with an outstretched hand in search of investors who agree to finance the development of domestic technology of nanopore sequencing.
The very first and most popular nanion pore sequencer MinION (the minimum set in the UK is $ 1000, in Russia - 150 thousand rubles), developed by Oxford Nanopore Technologies (ONT), operates on disposable cells, each of which costs $ 900 (in Russia - 135 thousand rubles). Such cells allow you to digitize DNA 10 ... 20 Gb in 2 ... 3 days. This is too little for sequencing the human genome (you need> 100 Gb), but too much for all other clinical diagnostic purposes (<1 Gb is enough).
Recently started selling more productive (and more expensive) models - PromethION 24/48 ($ 165,000 / $ 285,000), aimed primarily at full-genome in-line sequencing. More expensive are their more efficient consumables ($ 2,000 / unit,> 100 Gb). True, they can be cheaper in bulk ($ 1,800,000 for 2,880 pcs., $ 625 / pcs.), But such acquisitions (200 ... 250 million rubles per batch) are not permissible for the Russian budget.
And not only for Russian. Therefore, the most attention of visitors to the recent ONT conference (May 22-24) was attracted by the announcement of the start of sales of Flongle, an adapter insert for the MinION sequencer, which allows working with less efficient (~ 1 Gb), but relatively cheap ($ 90) disposable cells.

https://nanoporetech.com/products
The appearance of Flongle can cause an explosive growth in the use of NGS (next generation sequencing) in clinical practice and displace conventional PCR diagnostics, whose market size in Russia exceeds one billion rubles, and in the world it is measured in billions of dollars. But for most Russian patients, NGS diagnostics will still remain an inadmissible luxury, since the cost of cells for Flongle in the domestic market will be more than 10 thousand rubles. And if we add to it the cost of expendable reagents, transport costs and overhead, then the estimated cost of sequencing one DNA / RNA sample will not fall below 20 thousand rubles.
This implies the need to develop domestic nanopore sequencers and provide them with cheap consumables and reagents.
Nanorider
Significantly reduce the cost of nanopore sequencing is possible by reusing cells for Flongle and import substitution of the necessary reagents for working with them. But first you need to get a test device that allows you to monitor the formation of bilayer lipid membranes (BLM) in 126 wells of the working cell, and evaluate the quantity and quality of nanopores embedded in these membranes. Such a device ("Nanorider"?) Should track changes in the electrical conductivity (and / or impedance) of individual holes during the formation of BLMs on them, and register picoamperous currents flowing through single ion channels.
It is possible to compose a reader connected to the computer via USB 2.0 from standard elements: FPGA (100 ... 200 $), current amplifiers (MAX9923FEUB - 200 ... 300 rubles), ADC (1 ... 2 thousand rubles), etc. ... It’s necessary to develop only a board with a non-standard contact pad (10x13) type LGA (Land Grid Array) for docking with a cell from Flongle. Or with its simplified domestic counterpart containing a smaller number (16 ... 32) of sensor wells.

Touch Pad Cell Flongle
Reducing the design productivity of cells by 4 ... 8 times (up to 10 ... 20 Mb) is not critical for solving many problems (determining pathogens, HLA typing, establishing paternity, identification, etc.), but it allows you to collect a nanopore sequencer on conventional amplifiers picoampere currents. And do without the use of special 512-channel chips, the development of which the Oxfordians have invested tens of millions of dollars. True, each multichannel nanoreader must contain more than a dozen current amplifiers + ADCs, but it can still cost several times less than the proprietary MinION + Flongle combination (> 200 thousand rubles). As for the low performance of the nanoreader, for Russian users who do not have decent computers (doctors, biologists, biohackers, etc.), it can even be useful.
Kilorider
The 512-channel picoampere current amplifiers developed by the Oxfordians are contained in each disposable cell for the MinION. Waste cells can be thrown away, but it is better to use the chips contained in them to produce Flongle analogues that are characterized by increased productivity. If these chips are equipped with surface contacts, they will be able to work with contact cells containing 2048 (512x4) sensor wells.
Designing cells with such LGAs will not be easy, but you need to take into account that the contact requirements here are less stringent than processors and motherboards, in which one bad contact can disable the entire system. For a sequencer, the inoperability of even half of the cell contacts can be considered acceptable. The remaining half is enough to ensure performance at the level of 5 ... 10 Gb.
The logical development of this idea will be the combination of several similar readers in one device. This will allow you to get an analog of the GridION X5 sequencer, designed for simultaneous work with five (or more) reusable cells of domestic production. Such a sequencer may be in demand in many clinical diagnostic laboratories.

https://store.nanoporetech.com/devices
Disposable PromethION sequencer cells contain 9,000 sensor wells on chips. And the chips themselves are 3000-channel amplifiers of picoampere currents. The stream of information that they give out with 24 (PromethION 24) or 48 (PromethION 48) information cells is so large that in a raw form it cannot be transferred to a regular external computer. Therefore, sequencers of this type must be equipped with their own supercomputer. But if the sequencer will work only with one cell, then it can be connected to a regular gaming laptop with a Thunderbolt 3.0 port.
Alteration of free (ejected) chips from ONT to work with contact cells is not an insoluble problem. But sequencers built on the use of such chips can only work with ONT software. And the efforts that will have to be spent on remaking these programs can negate the benefits of using off-the-shelf chips. And developing such chips from scratch is too expensive.
Mega Rider
This year, beta testing of the nanopore sequencer from AXBIO (USA) is due to begin. It uses chips with a million channels of current amplification produced in Japan by the Israeli company Tower Semiconductor Ltd ... Consumables and the sale of these sequencers will be handled by the Chinese branch of AXBIO.

In the United States and the European Union, more than 7,000 patents are devoted to nanopore sequencing, which creates unpredictable financial risks for local companies trying to deal with such technologies. Apparently, this is the main reason for such a high internationalization of this development. And it allows Russian companies and laboratories to participate without any restrictions in testing and improving AXBIO technology, as well as producing (or collecting) such sequencers on their territory.
Terabytes of raw data obtained by a nanopore sequencer with megapixel chips (mega reader), it is desirable to filter. And transfer to the computer only the longest and highest-quality reads. This will not only improve the quality of sequencing, but also reduce the requirements for the computer, which is also important for mass production and widespread use of genomic sequencers. In addition, the excessive productivity of mega readers can reduce the duration of sequencing of the human genome from 2 ... 3 days (for ONT) to several hours.
Technology Sum
ONT's success was facilitated by the organization of parallel pipeline optimization of the five main elements of nanopore sequencing technology — membranes, nanopores, “motor” (helicase), sequencing conditions, and decoding algorithms for read signals.

https://nanoporetech.com/resource-centre/videos/sub1000
The guaranteed shelf life of cells for nanopore sequencing depends mainly on the stability of the bilayer lipid membrane. Now it is 6 weeks, although usually such membranes are destroyed in a few days. To increase their shelf life, hydrophobic polymer additives, surface-active block copolymers, cholesterol, antioxidants, cross-linking of fatty acids, etc. are used, and such modifications can interfere with the formation of ion channels. Therefore, it is very difficult to achieve BLM stability.
Radically solving this problem allows the formation of membranes in the working cells of the sequencer and the introduction of nanopores into them immediately before use. Self-preparation of cells for work will provide the opportunity for their regeneration and reuse. As for the shelf life, for dry cells it is almost unlimited, and the reagents necessary for their activation and regeneration can be stored for months.
The bottleneck in the technology of nanowire sequencing is at the same time the bottleneck of the nanopore, on which the quality of the read information depends. Therefore, one of the main tasks of ONT was the search for a membrane protein, which allows obtaining results of acceptable quality.
The first such protein was CsgG, an outer membrane protein found in many gram-negative bacteria. Nine of its subunits form an ion channel in the membrane with wide entrance and exit vestibules, and one bottleneck. But it was possible to obtain sufficiently high-quality nanopores (R9, R9.4.1, R9.5.1) only after enumerating and analyzing hundreds of modified CsgG variants. This summer, cells with the next generation pores (R10 or R10b) should appear on sale, featuring increased accuracy in reading homopolymer repeats.

https://nanoporetech.com/about-us/news/london-calling-clive-brown-and-team-plenary
ONT is an essential component of ONT's nanopore sequencing technology, a helicase, a molecular motor that unwinds double-stranded DNA and inhibits the progress of single-stranded DNA through a nanopore. The maximum speed of DNA unwinding with such a motor now reaches 450 nucleotides per second, although the speed of MinION electronic components allows it to be increased to 1000 nucleotides per second. Theoretically, accelerating the work of helicase can double the performance of MinION, but in practice this can lead to poor sequencing. Therefore, faster and more efficient helicases are unlikely to improve the performance of nanopore sequencers such as MinION and PromethION.
The quality of sequencing depends on many conditions - the magnitude of the voltage applied to the membrane, the electrical conductivity and the ionic composition of the reaction mixture, the amount and nature of impurities clogging the pores, the ability to clean clogged pores by changing the polarity of the potential, the structure of the adapters used, etc., etc. ... Therefore, nanopore sequencing requires careful optimization of all its parameters and compositions of working solutions (Run Conditions), which can adversely affect the quality of digitization of nucleotide sequences awns. For example, it was recently shown that a decrease in the concentration of ATP cleaved by helicase gradually decreases the accuracy of sequencing results, and now Oxfordians are trying to eliminate this problem by introducing an ATP regeneration system into the reaction mixture.
The fifth element of nanopore sequencing technology is artificial intelligence built on recurrent neural networks (RNN) and on the use of some special algorithms and programs that improve the recognition of homogeneous repeats (Flip-flop, Medaka). In July, the next release of the Guppi and MinKNOW programs should appear, in which the Oxfordians promise to add the ability to determine methylated bases (5mC and 6mA). This is important not only for epigenomic studies, but also to improve the recognition of conventional bases.
The large volumes of information read by nanopore sequencers and the complexity of its processing impose increased requirements on the hardware of such software and hardware systems. Therefore, not every computer is suitable for connecting MinION, the latest generation of graphics accelerators are installed in GridION and PromethION, and Oxford Nanopore Technologies became one of the first buyers of a large batch of Jetson AGX Xavier (NVIDIA) modules designed for devices with artificial intelligence.
More recently, nanopore technology was considered to be nothing more than a useful complement to more accurate fluorescence and semiconductor sequencing technologies. But a gradual increase in the accuracy of reading DNA, combined with a large length of readable sequences, allowed it, if not to get ahead of all competitors, then at least determine the main vector of development of genomic sequencing technologies.

It may take quite a while to master all the elements of nanopore sequencing technology, but this is not the biggest problem. The main problem is the lack of targeted funding for such integrated developments in Russia. True, the recently approved Federal Scientific and Technical Program for the Development of Genetic Technologies for 2019–2027 (section "Program Implementation Directions") says that in the short term (3 ... 6 years) a prototype device for high-performance genomic sequencing will be developed. But finding this single line on 30 pages of text is not so easy. Easier to negotiate with the Chinese on the purchase of AXBIO sequencers.
Even simpler is to find craftsmen who can assemble the simplest nanorider “on the knee”. And then walk with an outstretched hand in search of investors who agree to finance the development of domestic technology of nanopore sequencing.
