What's inside XGBoost, and where does Go
In the world of machine learning, one of the most popular types of models is a decisive tree and ensembles based on them. The advantages of trees are: simplicity of interpretation, no restrictions on the type of initial dependence, soft requirements for the sample size. Trees have a major drawback - a tendency to retrain. Therefore, almost always trees are combined into ensembles: a random forest, gradient boosting, etc. The complex theoretical and practical tasks are to compile trees and combine them into ensembles.
In this article, we will consider the procedure for the formation of predictions from the already trained model of an ensemble of trees, the features of implementations in popular gradient boosting libraries
Consider first the general provisions. Usually work with trees, where:
I took this illustration from the XGBoost documentation.
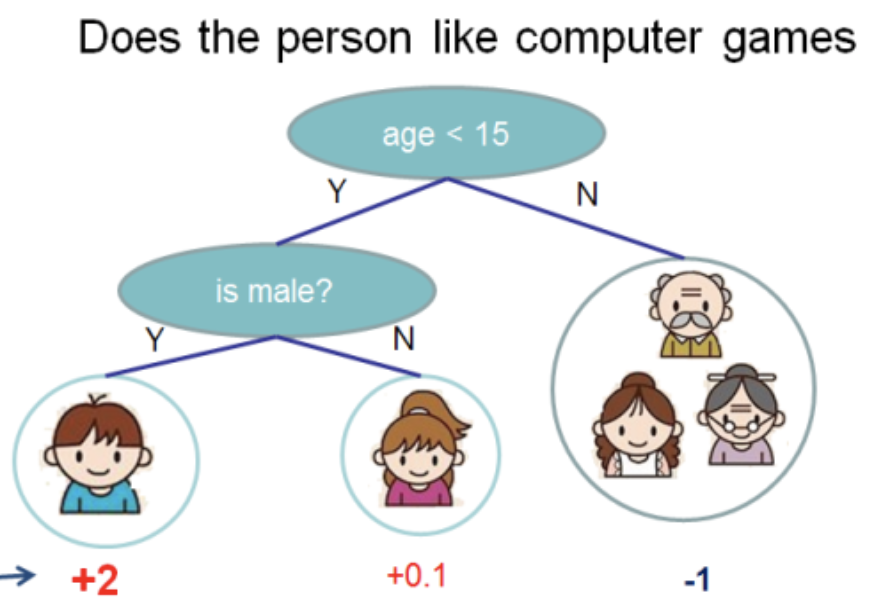
In this tree we have 2 nodes, 2 decision rules and 3 sheets. Below the circles are the values - the result of applying the tree to an object. Usually, a transformation function is applied to the result of the calculation of a tree or ensemble of trees. For example, sigmoid for a binary classification problem.
To obtain predictions from an ensemble of trees obtained by the gradient boosting method, you need to add the prediction results of all the trees:
Hereinafter, the code will be on
Next, consider what is hidden in
In
The decision rule is implemented in the function
Two things follow from here:
Central place is the class
The following features can be distinguished:
In order to be able to trace the path from the method call
In the function,
LightGBM has no node data structure. Instead, the tree data structure
Consider the decision rule for a categorical trait:
It can be seen that, depending on the
i.e., for example, for a categorical attribute, it is
This approach has one major drawback: if the categorical feature can take large values, for example 100,500, then a bit field of up to 12,564 bytes in size will be created for each decision rule!
Therefore, it is desirable to renumber the values of the categorical attributes so that they go continuously from 0 to the maximum value .
For my part, I made explanatory changes to
Working with real signs is not much different from
For writing a loaded backend service, a popular language is
Therefore, I decided to write a library in pure

The main features of the library:
I will give here a minimal program on
The library API is minimalistic. To use the model,
Despite the fact that the language
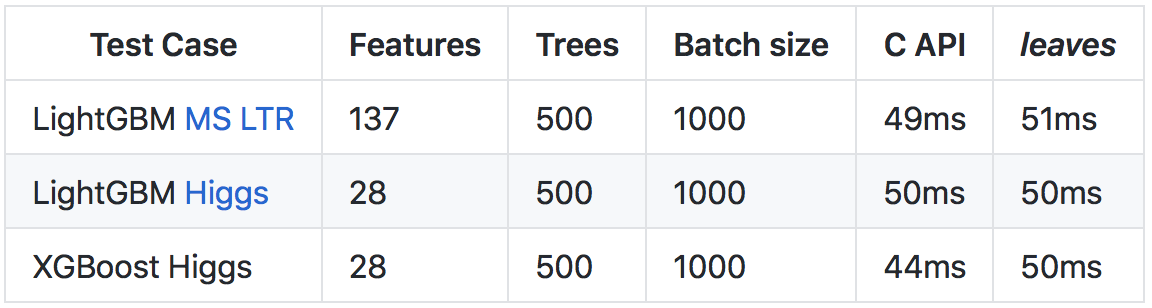
More details about the results and the way of comparisons are in the repository on github .
I hope this article I opened the door in the implementation of trees in libraries
For those who are interested in the topic of using gradient boosting models in their Go services, I recommend reading the leaves library . With the help,
Successes!
In this article, we will consider the procedure for the formation of predictions from the already trained model of an ensemble of trees, the features of implementations in popular gradient boosting libraries
XGBoostand LightGBM. As well as the reader will get acquainted with the libraryleaves for Go, which allows you to make predictions for ensembles of trees, without using the C API of the original libraries.Where do trees grow from?
Consider first the general provisions. Usually work with trees, where:
- splitting in a node occurs by one attribute
- tree is binary - each node has a left and right descendant
- in the case of a real feature, the decision rule consists of comparing the value of the feature with the threshold value
I took this illustration from the XGBoost documentation.

In this tree we have 2 nodes, 2 decision rules and 3 sheets. Below the circles are the values - the result of applying the tree to an object. Usually, a transformation function is applied to the result of the calculation of a tree or ensemble of trees. For example, sigmoid for a binary classification problem.
To obtain predictions from an ensemble of trees obtained by the gradient boosting method, you need to add the prediction results of all the trees:
double pred = 0.0;
for (auto& tree: trees)
pred += tree->Predict(feature_values);
Hereinafter, the code will be on
C++, since exactly in this language are written XGBoostand LightGBM. I will omit irrelevant details and try to give the most concise code. Next, consider what is hidden in
Predict, and how the tree data structure is organized.XGBoost trees
In
XGBoostthere are several classes (in the sense of OOP) trees. We will talk about RegTree(see include/xgboost/tree_model.h), which, according to the documentation, is the main one. If you leave only the details that are important for predictions, the members of the class look as simple as possible:classRegTree {// vector of nodesstd::vector<Node> nodes_;
};
The decision rule is implemented in the function
GetNext. The code is slightly modified, without affecting the result of the calculations:// get next position of the tree given current pidint RegTree::GetNext(int pid, float fvalue, bool is_unknown) const {
constauto& node = nodes_[pid]
float split_value = node.info_.split_cond;
if (is_unknown) {
return node.DefaultLeft() ? node.cleft_ : node.cright_;
} else {
if (fvalue < split_value) {
return node.cleft_;
} else {
return node.cright_;
}
}
}
Two things follow from here:
RegTreeworks only with real signs (typefloat)- missing attribute values supported
Central place is the class
Node. It contains the local tree structure, the decision rule and the leaf value:classNode {public:
// feature index of split conditionunsignedSplitIndex()const{
return sindex_ & ((1U << 31) - 1U);
}
// when feature is unknown, whether goes to left childboolDefaultLeft()const{
return (sindex_ >> 31) != 0;
}
// whether current node is leaf nodeboolIsLeaf()const{
return cleft_ == -1;
}
private:
// in leaf node, we have weights, in non-leaf nodes, we have split conditionunion Info {
float leaf_value;
float split_cond;
} info_;
// pointer to left, rightint cleft_, cright_;
// split feature index, left split or right split depends on the highest bitunsigned sindex_{0};
};
The following features can be distinguished:
- sheets are presented as nodes that have
cleft_ = -1 - the field
info_is represented asunion, i.e. two types of data (in this case, the same) divide one section of memory depending on the type of node - the most significant bit in
sindex_is responsible for where the object goes down for which the value of the attribute is omitted
In order to be able to trace the path from the method call
RegTree::Predictto the receipt of the answer, I’ll give the missing two functions:float RegTree::Predict(const RegTree::FVec& feat, unsigned root_id) const {
int pid = this->GetLeafIndex(feat, root_id);
return nodes_[pid].leaf_value;
}
int RegTree::GetLeafIndex(const RegTree::FVec& feat, unsigned root_id) const {
auto pid = static_cast<int>(root_id);
while (!nodes_[pid].IsLeaf()) {
unsigned split_index = nodes_[pid].SplitIndex();
pid = this->GetNext(pid, feat.Fvalue(split_index), feat.IsMissing(split_index));
}
return pid;
}
In the function,
GetLeafIndexwe cycle down the nodes of the tree until we get to the leaf.LightGBM trees
LightGBM has no node data structure. Instead, the tree data structure
Tree(file include/LightGBM/tree.h) contains arrays of values, where the index is the node number. The values in the leaves are also stored in separate arrays.classTree {// Number of current leavesint num_leaves_;
// A non-leaf node's left childstd::vector<int> left_child_;
// A non-leaf node's right childstd::vector<int> right_child_;
// A non-leaf node's split feature, the original indexstd::vector<int> split_feature_;
//A non-leaf node's split threshold in feature valuestd::vector<double> threshold_;
std::vector<int> cat_boundaries_;
std::vector<uint32_t> cat_threshold_;
// Store the information for categorical feature handle and mising value handle.std::vector<int8_t> decision_type_;
// Output of leavesstd::vector<double> leaf_value_;
};
LightGBMsupports categorical features. Support is provided using a bitfield that is stored in cat_threshold_for all nodes. In cat_boundaries_stores, to which node which part of the bit field corresponds. The field threshold_for the categorical case is translated into intand corresponds to the index in cat_boundaries_to search for the beginning of the bit field. Consider the decision rule for a categorical trait:
intCategoricalDecision(double fval, int node)const{
uint8_t missing_type = GetMissingType(decision_type_[node]);
int int_fval = static_cast<int>(fval);
if (int_fval < 0) {
return right_child_[node];;
} elseif (std::isnan(fval)) {
// NaN is always in the rightif (missing_type == 2) {
return right_child_[node];
}
int_fval = 0;
}
int cat_idx = static_cast<int>(threshold_[node]);
if (FindInBitset(cat_threshold_.data() + cat_boundaries_[cat_idx],
cat_boundaries_[cat_idx + 1] - cat_boundaries_[cat_idx], int_fval)) {
return left_child_[node];
}
return right_child_[node];
}
It can be seen that, depending on the
missing_typevalue, it NaNautomatically lowers the solution along the right branch of the tree. Otherwise, it NaNis replaced by 0. Finding a value in a bit field is quite simple:boolFindInBitset(constuint32_t* bits, int n, int pos){
int i1 = pos / 32;
if (i1 >= n) {
returnfalse;
}
int i2 = pos % 32;
return (bits[i1] >> i2) & 1;
}
i.e., for example, for a categorical attribute, it is
int_fval=42checked whether the 41st (numbering with 0) bits are set in the array. This approach has one major drawback: if the categorical feature can take large values, for example 100,500, then a bit field of up to 12,564 bytes in size will be created for each decision rule!
Therefore, it is desirable to renumber the values of the categorical attributes so that they go continuously from 0 to the maximum value .
For my part, I made explanatory changes to
LightGBMand accepted them . Working with real signs is not much different from
XGBoost, and I will skip this for short.leaves - Go library for predictions
XGBoostand LightGBMvery powerful libraries for building gradient boost models on decision trees. To use them in the backend service, where machine learning algorithms are necessary, it is necessary to solve the following tasks:- Periodic training of models in offline
- Delivery models in the backend service
- Online Model Survey
For writing a loaded backend service, a popular language is
Go. To drag XGBoostor LightGBMthrough C API and cgo is not the easiest solution - the program builds up, because of careless handling you can catch SIGTERMproblems with the number of system threads (OpenMP inside libraries vs go runtime threads). Therefore, I decided to write a library in pure
Goprediction using models built in XGBoostor LightGBM. It is called leaves. 
The main features of the library:
- For
LightGBMmodels- Reading models from a standard format (text)
- Support for real and categorical features
- Missing Value Support
- Optimization of work with categorical variables
- Optimizing predictions with predictive-only data structures
- For
XGBoostmodels- Reading models from a standard format (binary)
- Missing Value Support
- Prediction optimization
I will give here a minimal program on
Gowhich loads the model from the disk and displays the prediction on the screen:package main
import(
"bufio""fmt""os""github.com/dmitryikh/leaves"
)
func main(){
// 1. Открываем файл с моделью
path := "lightgbm_model.txt"
reader, err := os.Open(path)
if err != nil {
panic(err)
}
defer reader.Close()
// 2. Читаем модель LightGBM
model, err := leaves.LGEnsembleFromReader(bufio.NewReader(reader))
if err != nil {
panic(err)
}
// 3. Делаем предсказание!
fvals := []float64{1.0, 2.0, 3.0}
p := model.Predict(fvals, 0)
fmt.Printf("Prediction for %v: %f\n", fvals, p)
}
The library API is minimalistic. To use the model,
XGBoostjust call the method leaves.XGEnsembleFromReader, instead of the one above. Predictions can be made in batches by calling methods PredictDenseor model.PredictCSR. More usage scenarios can be found in the tests for leaves . Despite the fact that the language
Gois slower C++(mainly due to the heavier runtime and runtime checks), due to a number of optimizations, it was possible to achieve a prediction speed comparable to the C API call of the original libraries. 
More details about the results and the way of comparisons are in the repository on github .
Behold the root
I hope this article I opened the door in the implementation of trees in libraries
XGBoostand LightGBM. As you can see, the basic constructs are fairly simple, and I encourage readers to take advantage of open source — to study the code when there are questions about how it works. For those who are interested in the topic of using gradient boosting models in their Go services, I recommend reading the leaves library . With the help,
leavesyou can quite simply use the leading edge solutions in machine learning in your production environment, practically not losing in speed compared to the original C ++ implementations. Successes!
