R + C + CUDA = ...
 Sometimes there is a need to speed up the calculation, and it is desirable immediately at times. At the same time, you have to give up convenient, but slow tools and resort to something lower-level and faster. R has fairly advanced capabilities for working with dynamic libraries written in C / C ++, Fortran, or even Java. Out of habit, I prefer C / C ++.
Sometimes there is a need to speed up the calculation, and it is desirable immediately at times. At the same time, you have to give up convenient, but slow tools and resort to something lower-level and faster. R has fairly advanced capabilities for working with dynamic libraries written in C / C ++, Fortran, or even Java. Out of habit, I prefer C / C ++.R and C
Immediately make a reservation that I work under Debian Wheezy (under Windows, there are probably some nuances). When writing a library in C for R, consider the following:
- Functions written in C and called in R must be of type void. This means that if a function returns any results, then they must be returned through the function arguments
- All arguments are passed by reference (and when working with pointers, you must not lose vigilance!)
- In C, it is desirable to include the code
R.handRmath.h(if the mathematical functions R are used)
Let's start with a simple function that calculates the scalar product of two vectors:
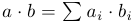
#include
void iprod(double *v1, double *v2, int *n, double *s) {
double ret = 0;
int len = *n;
for (int i = 0; i < len; i++) {
ret += v1[i] * v2[i];
}
*s = ret;
}
Next, you need to get a dynamic library - you can directly through gcc, or you can use this command (by the way, you should remember the output, because it will be useful to us in the future):
R CMD SHLIB inner_prod.c
At the output, we get the inner_prod.so file, for loading of which we use the function
dyn.load(). To call the function itself in C, we use it .C()(there is .Call()also .External(), but with a slightly different functionality, and sometimes there is heated debate between the supporters of .C () and .Call () ). I only note that when writing code in C for calling through it it turns out to be cleaner and more readable. Particular attention should be paid to the correspondence of types of variables in R and C ( this is described in detail in the documentation for the function ). Wrapper Function on R:.C().C()iprodс <- function(a, b) {
if (!is.loaded('iprod')) {
dyn.load("inner_prod.so")
}
n <- length(a)
v <- 0
return(.C("iprod", as.double(a), as.double(b), as.integer(n), as.double(v))[[4]])
}
Now you can find out what we have achieved:
> n <- 1e7; a <- rnorm(n); b <- rnorm(n);
> iprodс(a, b)
[1] 3482.183
And a little check:
> sum(a * b)
[1] 3482.183
In any case, he thinks right.
R and CUDA
To take advantage of all the benefits that Nvidia Graphics Accelerator provides in Debian, you need to have the Nvidia proprietary driver and package installed
nvidia-cuda-toolkit. CUDA certainly deserves a separate huge topic, and since my level in this topic is “newbie”, I will not scare people with my crooked and incomplete code, but let me copy a few lines from the manuals. Suppose it is necessary to raise each element of the vector to the third power and find the Euclidean norm of the resulting vector:

To make it easier to work with the GPU, we use a library of parallel algorithms
Thrust, which allows us to abstract from low-level CUDA / C operations. In this case, the data is presented in the form of vectors to which some standard algorithms are applied (elementwise operations, reductions, prefix-sums, sorting).#include
#include
#include
// Функтор, выполняющий возведение числа в 6 степень на GPU (device)
template
struct power{
__device__
T operator()(const T& x) const{
return std::pow(x, 6);
}
};
extern "C" void nrm(double *v, int *n, double *vnorm) {
// Вектор, хранимый в памяти GPU, в который копируется содержимое *v
thrust::device_vector dv(v, v + *n);
// Reduce-трансформация вектора, т.е. сначала к каждому члену вектора применятся функтор power
// потом полученные числа складываются.
*vnorm = std::sqrt( thrust::transform_reduce(dv.begin(), dv.end(), power(), 0.0, thrust::plus()) );
}
Use is
extern "C"mandatory here, otherwise R will not see the nrm () function. We will now use nvcc to compile the code. Remember the output of the command R CMD SHLIB...? Here is a bit of it and adapt it so that a library using CUDA / Thrust can be called from R without problems:nvcc -g -G -O2 -arch sm_30 -I/usr/share/R/include -Xcompiler "-Wall -fpic" -c thr.cu thr.o
nvcc -shared -lm thr.o -o thr.so -L/usr/lib/R/lib -lR
At the output we get DSO thr.so. The wrapper function is practically no different:
gpunrm <- function(v) {
if (!is.loaded('nrm'))
dyn.load("thr.so")
n <- length(v)
vnorm <- 0
return(.C("nrm", as.double(v), as.integer(n), as.double(vnorm))[[3]])
}
The graph below clearly shows how the runtime grows depending on the length of the vector. It is worth noting that if simple operations such as addition / subtraction prevail in the calculations, then there will be practically no difference between the calculation time on the CPU and GPU. Moreover, it is very likely that the GPU will lose due to the overhead of working with memory.
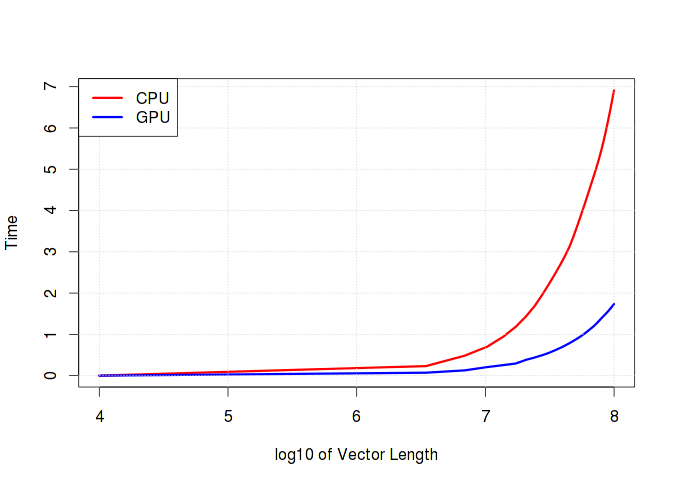
Hidden text
gpu_time <- c()
cpu_time <- c()
n <- seq.int(1e4, 1e8, length.out=30)
for (i in n) {
v <- rnorm(i, 1000)
gpu_time <- c(gpu_time, system.time({p1 <- gpunrm(v)})[1])
cpu_time <- c(cpu_time, system.time({p2 <- sqrt(sum(v^6))})[1])
}
Conclusion
In fact, in R operations for working with matrices and vectors are very well optimized, and the need to use a GPU in everyday life does not arise very often, but sometimes the GPU can significantly reduce the calculation time. CRAN already has ready-made packages (for example,
gputools) adapted for working with the GPU ( here you can read more about this).References
1. An Introduction to the .C Interface to R
2. Calling C Functions in R and Matlab
3. Writing CUDA C extensions for R
4. Thrust :: CUDA Toolkit Documentation
PS Corrected the first code fragment as recommended by vird .
