R Code Acceleration Strategies, Part 2
- Transfer
The for loop in R can be very slow if it is applied in its pure form, without optimization, especially when dealing with large data sets. There are a number of ways to make your code faster, and you will probably be surprised to find out how much.
This article describes several approaches, including simple changes in logic, parallel processing and
Let's try to speed up the code with a for loop and a conditional statement (if-else) to create a column that is added to the data set (data frame, df). The code below creates this initial dataset.
In the first part : vectorization, only true conditions, ifelse.
In this part: which, apply, byte compilation, Rcpp, data.table, results.
Using the command
We use a function

Using apply and the for loop in R
This is probably not the best example to illustrate the efficiency of byte compilation, since the resulting time is slightly higher than the usual form. However, for more complex functions, byte compilation has proven effective. I think it's worth a try on occasion.

Apply, for loop and byte code compilation
Let's get to a new level. Prior to this, we increased speed and productivity through various strategies and found that use was
Below is the same logic implemented in C ++ using the Rcpp package. Save the code below as “MyFunc.cpp” in your R session working directory (or you will have to use sourceCpp using the full path). Please note that a comment is
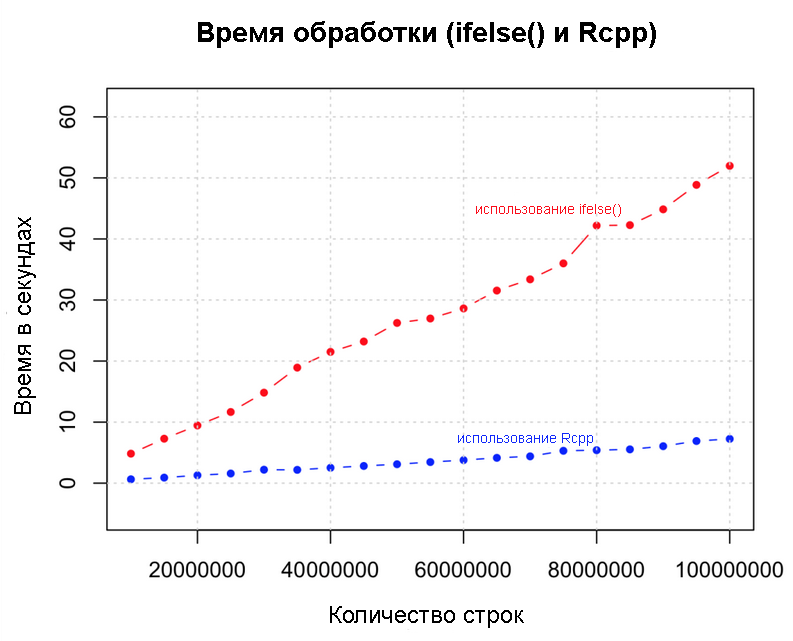
Performance
Parallel processing:
Remove more unnecessary objects in your code using
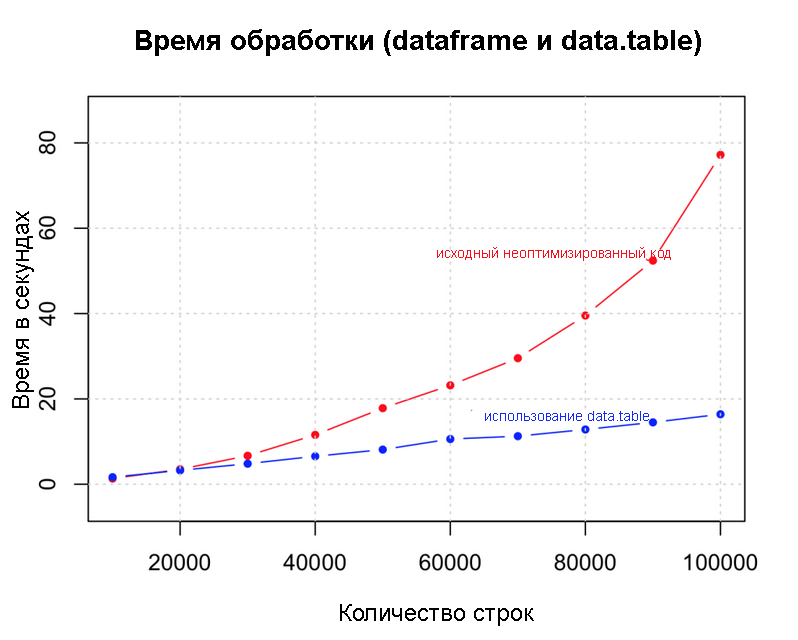
Dataframe and data.table
Method: Speed, number of lines in df / elapsed time = n lines per second
Original: 1X, 120,000 / 140.15 = 856.2255 lines per second (normalized to 1)
Vectorized: 738X, 120,000 / 0.19 = 631578.9 lines per second
Only true conditions: 1002X , 120000 / 0.14 = 857142.9 lines per second
ifelse: 1752X, 1200000 / 0.78 = 1500000 lines per second
which: 8806X, 2985984 / 0.396 = 7540364 lines per second
Rcpp: 13476X, 1200000 / 0.09 = 11538462 lines per second
The numbers above approximate and based on random starts. No calculation results for
This article describes several approaches, including simple changes in logic, parallel processing and
Rcpp, increasing the speed by several orders of magnitude, so that it will be possible to normally process 100 million rows of data or even more. Let's try to speed up the code with a for loop and a conditional statement (if-else) to create a column that is added to the data set (data frame, df). The code below creates this initial dataset.
# Создание набора данных
col1 <- runif (12^5, 0, 2)
col2 <- rnorm (12^5, 0, 2)
col3 <- rpois (12^5, 3)
col4 <- rchisq (12^5, 2)
df <- data.frame (col1, col2, col3, col4)
In the first part : vectorization, only true conditions, ifelse.
In this part: which, apply, byte compilation, Rcpp, data.table, results.
Using which ()
Using the command
which()to select the rows, one third of the speed can be achieved Rcpp.# Спасибо Гейб Бекер
system.time({
want = which(rowSums(df) > 4)
output = rep("less than 4", times = nrow(df))
output[want] = "greater than 4"
})
# количество строк = 3 миллиона (примерно)
user system elapsed
0.396 0.074 0.481
Use the apply function family instead of for loops
We use a function
apply()to implement the same logic and compare it with a vectorized for loop. The results grow with an increase in the number of orders, but they are slower than the ifelse()versions where the verification was done outside the loop. This may be useful, but it may take some ingenuity for complex business logic.# семейство apply
system.time({
myfunc <- function(x) {
if ((x['col1'] + x['col2'] + x['col3'] + x['col4']) > 4) {
"greater_than_4"
} else {
"lesser_than_4"
}
}
output <- apply(df[, c(1:4)], 1, FUN=myfunc) # применить 'myfunc' к каждой строке
df$output <- output
})

Using apply and the for loop in R
Use byte compilation for cmpfun () functions from compiler package instead of function itself
This is probably not the best example to illustrate the efficiency of byte compilation, since the resulting time is slightly higher than the usual form. However, for more complex functions, byte compilation has proven effective. I think it's worth a try on occasion.
# побайтовая компиляция кода
library(compiler)
myFuncCmp <- cmpfun(myfunc)
system.time({
output <- apply(df[, c (1:4)], 1, FUN=myFuncCmp)
})

Apply, for loop and byte code compilation
Use rcpp
Let's get to a new level. Prior to this, we increased speed and productivity through various strategies and found that use was
ifelse()most effective. What if we add another zero? Below we implement the same logic with Rcpp, with a data set of 100 million rows. We will compare speeds Rcppand ifelse().library(Rcpp)
sourceCpp("MyFunc.cpp")
system.time (output <- myFunc(df)) # функция Rcpp ниже
Below is the same logic implemented in C ++ using the Rcpp package. Save the code below as “MyFunc.cpp” in your R session working directory (or you will have to use sourceCpp using the full path). Please note that a comment is
// [[Rcpp::export]]required and must be placed immediately before the function that you want to execute from R.// Источник для MyFunc.cpp
#include
using namespace Rcpp;
// [[Rcpp::export]]
CharacterVector myFunc(DataFrame x) {
NumericVector col1 = as(x["col1"]);
NumericVector col2 = as(x["col2"]);
NumericVector col3 = as(x["col3"]);
NumericVector col4 = as(x["col4"]);
int n = col1.size();
CharacterVector out(n);
for (int i=0; i 4){
out[i] = "greater_than_4";
} else {
out[i] = "lesser_than_4";
}
}
return out;
}

Performance
RcppandifelseUse parallel processing if you have a multi-core computer
Parallel processing:
# параллельная обработка
library(foreach)
library(doSNOW)
cl <- makeCluster(4, type="SOCK") # for 4 cores machine
registerDoSNOW (cl)
condition <- (df$col1 + df$col2 + df$col3 + df$col4) > 4
# параллелизация с векторизацией
system.time({
output <- foreach(i = 1:nrow(df), .combine=c) %dopar% {
if (condition[i]) {
return("greater_than_4")
} else {
return("lesser_than_4")
}
}
})
df$output <- output
Delete variables and clear memory as early as possible
Remove more unnecessary objects in your code using
rm()as soon as possible, especially before long loops. Sometimes it can help gc()at the end of each iteration of the loop.Use data structures that take up less memory
Data.table()Is a great example because it does not overload memory. This speeds up operations like data federation.dt <- data.table(df) # создать data.table
system.time({
for (i in 1:nrow (dt)) {
if ((dt[i, col1] + dt[i, col2] + dt[i, col3] + dt[i, col4]) > 4) {
dt[i, col5:="greater_than_4"] # присвоить значение в 5-й колонке
} else {
dt[i, col5:="lesser_than_4"] # присвоить значение в 5-й колонке
}
}
})

Dataframe and data.table
Speed: Results
Method: Speed, number of lines in df / elapsed time = n lines per second
Original: 1X, 120,000 / 140.15 = 856.2255 lines per second (normalized to 1)
Vectorized: 738X, 120,000 / 0.19 = 631578.9 lines per second
Only true conditions: 1002X , 120000 / 0.14 = 857142.9 lines per second
ifelse: 1752X, 1200000 / 0.78 = 1500000 lines per second
which: 8806X, 2985984 / 0.396 = 7540364 lines per second
Rcpp: 13476X, 1200000 / 0.09 = 11538462 lines per second
The numbers above approximate and based on random starts. No calculation results for
data.table(), byte code compilation and parallelization, because they will be very different in each case and depending on how you use them.